
RESEARCH
PROTEIN ENGINEERING
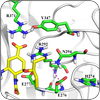
This research program uses simulation techniques deployed on DoD high-performance computing assets to realistically model the behavior of proteins and peptides in solvent solutions, investigate protein-protein interactions, and determine how mutations of specific amino acids affect the biological function of proteins. The program covers a broad range of topics, including issues of protein stability, generation of representative protein conformations, and selection of RNA aptamers.
Publications
Liu, J., G. J. Tawa, and A. Wallqvist. Identifying cytochrome P450 functional networks and their allosteric regulatory elements. PLOS ONE. 2013 December 3; 8(12):e81980. [PDF, PubMed]
Yeh, I.-C., D. R. Ripoll, and A. Wallqvist. Free energy difference in indolicidin attraction to eukaryotic and prokaryotic model cell membranes. Journal of Physical Chemistry B. 2012 March 15; 116(10):3387-3396. [PDF, PubMed]
Chaudhury, S., M. A. Olson, G. Tawa, A. Wallqvist, and M. S. Lee. Efficient conformational sampling in explicit solvent using a hybrid replica exchange molecular dynamics method. Journal of Chemical Theory and Computation. 2012 February; 8(2):677-687. [PDF, PubMed]
Koehler, J. W., L. C. Dupuy, A. R. Garrison, B. F. Beitzel, M. J. Richards, D. R. Ripoll, A. Wallqvist, S. Y. Teh, A. A. Vaewhongs, F. S. Vojdani, H. S. Padgett, and C. S. Schmaljohn. Novel plant-derived recombinant human interferons with broad spectrum antiviral activity. Antiviral Research. 2011 December; 92(3):461-469. [PDF, PubMed]
Olson, M. A., S. Chaudhury, and M. S. Lee. Comparison between self-guided Langevin dynamics and molecular dynamics simulations for structure refinement of protein loop conformations. Journal of Computational Chemistry. 2011 November 15; 32(14):3014-3022. [PDF, PubMed]
Hu, X., M. S. Lee, K. Kumar, and A. Wallqvist. An integrated docking pipeline for the prediction of large-scale protein-protein interactions. Proceedings of the HPCMP Users Group Conference. Schaumburg, IL. 2010 June 14-17; 228-232. [PDF, DTIC]
Hu, X., M. S. Lee, and A. Wallqvist. Interaction of the disordered Yersinia effector protein YopE with its cognate chaperone SycE. Biochemistry. 2009 December 1; 48(47):11158-11160. [PDF, PubMed]
Yeh, I.-C., and A. Wallqvist. Structure and dynamics of end-to-end loop formation of the penta-peptide Cys-Ala-Gly-Gln-Trp in implicit solvents. Journal of Physical Chemistry B. 2009 September 10; 113(36):12382-12390. [PDF, PubMed]
Chushak, Y., and M. O. Stone. In silico selection of RNA aptamers. Nucleic Acids Research. 2009 July; 37(12):e87. [PDF, PubMed]
Harbaugh, S., N. Kelley-Loughnane, M. Davidson, L. Narayanan, S. Trott, Y. G. Chushak, and M. O. Stone. FRET-based optical assay for monitoring riboswitch activation. Biomacromolecules. 2009 May 11; 10(5):1055-1060. [PDF, PubMed]
Bondugula, R., M. S. Lee, and A. Wallqvist. FIEFDom: a transparent domain boundary recognition system using a fuzzy mean operator. Nucleic Acids Research. 2009 February; 37(2):452-462. [PDF, PubMed]
Yeh, I.-C., M. S. Lee, and M. A. Olson. Calculation of protein heat capacity from replica-exchange molecular dynamics simulations with different implicit solvent models. Journal of Physical Chemistry B. 2008 November 27; 112(47):15064-15073. [PDF, PubMed]
Harbaugh, S. V., M. E. Davidson, Y. G. Chushak, N. Kelley-Loughnane, and M. O. Stone. Riboswitch-based sensor in low optical background. Proceedings of the Nanobiosystems: Processing, Characterization, and Applications - International Society of Optical Engineering. San Diego, CA. 2008 August 12-14; 7040:1-9. [PDF, DTIC]